Pbac
Pbac serves as a specialized genomic resource for studying microbial communities in giant panda gastrointestinal systems. This curated database addresses the unique bamboo-digesting adaptations of endangered ursids through two major releases: Version 1 established the foundational collection with 2,684 metagenome-assembled genomes (MAGs) derived from hybrid Illumina-Nanopore sequencing, including 354 high-quality representatives that revealed significant novel microbial diversity. Version 2 substantially enhanced genomic quality through integrated culturomics and PacBio HiFi sequencing, expanding to 466 species-level references (84% increase) with 213 newly documented species and five novel genera. Validation shows >92% mapping efficiency for microbiome analyses. Researchers can leverage Pbac for diverse applications including metagenomic profiling, conservation genomics, and investigating host-microbe coevolution in bamboo-digesting ecosystems. The database provides both individual genome access and bulk downloads of pre-compiled Kraken2 databases for efficient analysis pipelines.




Data Resource
Provided resource in Pbac v2 database.
Recent News & Blogs

04
SEP
Long-Read Metagenomic Sequencing of a Pooled Fecal Sample from Giant Pandas (ONT & PacBio HiFi)
A pooled fecal sample from three giant pandas was sequenced using Oxford Nanopore Technologies (ONT) and PacBio HiFi platforms, and the datasets......
03
SEP
Analysis of Giant Panda Gut Microbial Metagenomic Data Using Kraken2 and PBac v2 Databases
We have pre-built Kraken2 (v2.1.2) and Bracken (v2.9) reference database packages based on the PBac v2 database, which can be downloaded from this......

03
SEP
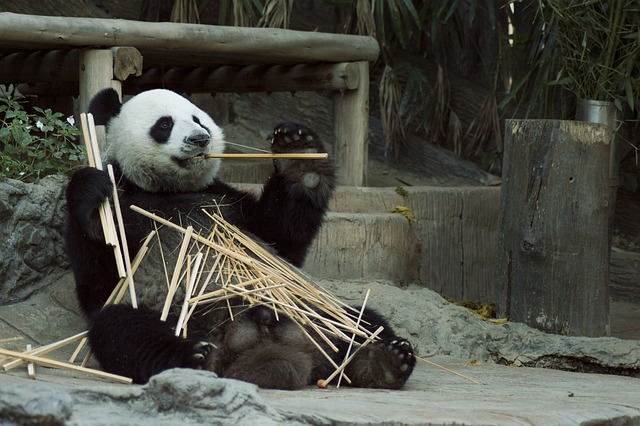
Taxonomic Relative Abundance Estimation via Kraken2 and Bracken
1. MAGs Generation Workflow2. Isolated Bacterial Genome Assembly Workflow3. Taxonomic Relative Abundance Estimation via Kraken2 and Bracken4.......
Copyright © 2025. bacbase.com All rights reserved. 蜀ICP备2023027719号



